Whether your palate runs to domestic or imported, a piece of cheese can be a real treat for the senses. Its smell, taste, and texture are all parts of its appeal. A big part of what makes that savory wonderfulness comes from the microbes in and on the cheese. Thanks to a team of researchers dedicated to studying those microbes, we have a better understanding of their importance to cheese and us.
A new study by researcher Rachel Dutton, of the University of California in San Diego, and her team expands on their earlier work into the microbial community present on the rind of cheese. It may seem like just the outer protective layer of cheese, but it is teeming with microbes that form an interactive and dynamic network.
Their new study was published June 23 in the journal mLife.
From Milk to a Slice of Heaven
From large scale professional cheese makers to home artisanal cheese crafters, the same basic processes apply. A starter culture of bacteria is added to milk from cows, goats, sheep or buffalo to form lactic acid from the milk sugar, lactose. Raw milk — milk that has not been pasteurized to kill disease-causing bacteria — can and is used to make cheese, but its consumers are at significant risk of developing a foodborne illness.
The starter bacteria determine what type of cheese will develop from the milk. The enzyme rennin coagulates the milk, separating the milk into curds and whey. Removal of the whey and compression of the curds — either manually or by machine — forms the shape. Fresh mozzarella or cheese curds are ready at this stage, but others go through brining and washing to reduce bacterial contamination, and aging.
Aging is a process key to 'ripening' the cheese and developing its full and distinctive flavor. And it's not as simple as storing the cheese and waiting a required time. Just like whiskey, the environment it ages in can make all the difference. While in the past, these aging environments were literally cheese caves, used because of their relatively stable temperature and humidity, today that term usually refers to a stable environment where cheese ripens. Temperatures between 45–58ºF, humidity of 80–98%, and fresh air make an environment suitable for aging cheese.
It's during this process that microbes get busy. The starter microbes die and other bacteria bloom, largely on the rind that develops. The microbial communities of cheese rinds include Firmicutes, Actinobacteria, Proteobacteria, and Bacteroidetes bacteria, and various species of fungus. The abundance and diversity of these microbes vary during the cheese ripening process, depending on the type of rind (bloomy from fungal growth, washed or natural) and whether the cheese maker wants to ripen it to soft, semi-hard, or hard.

Brie with bloomy rind

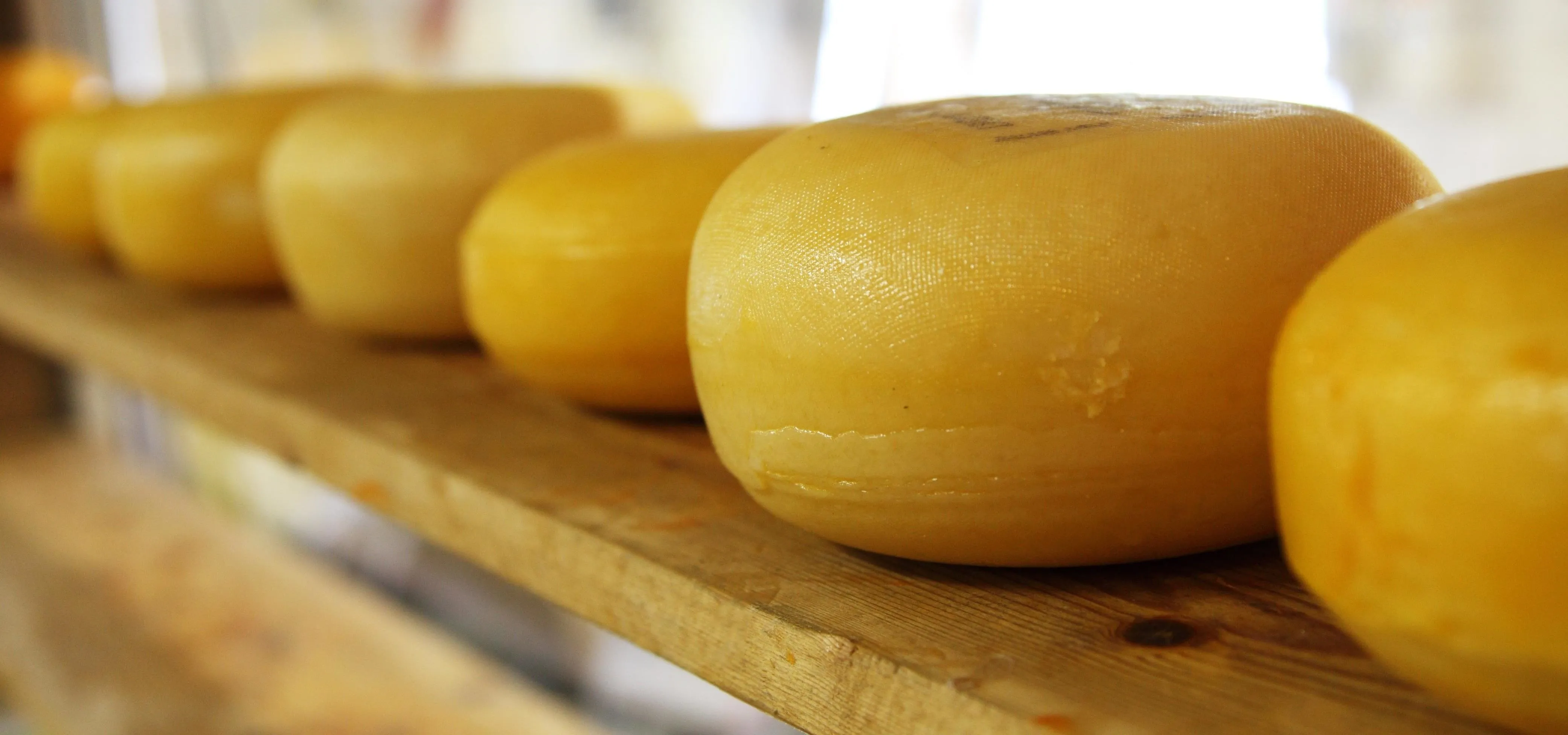
Hard cheese with rind

Brie with bloomy rind

Hard cheese with rind
Not only is the aging environment critical to developing the defining flavors of different cheeses, but it also supplies a source of bacterial acquisition and interaction that provides essential functions to the bacteria on the cheese rind.
Rind's Unappreciated Role
The rind that develops on cheese as it ages is a biofilm, a build-up of microbes that adhere to a surface. The biofilm that makes up cheese rind is a collection of microbes, including bacteria from the cheese itself, the aging environment, and the aging process that may involve inoculation with other fungi, as well as repeated washing of the rind by humans.
Three rind types result from different aging techniques.
Bloomy rinds like brie and Camembert are inoculated with fungi to create the dense, white rind. The microbes on brie break down proteins and fats on the rind and create volatile sulfur and ammonia compounds that provide the sharp odor of these cheeses. Natural rind cheeses may be cloth-wrapped, like St. Nectaire, and Tomme de Savoie cheddars. Washed rind cheeses, such as Taleggio, Gruyere, and Epoisses are repeatedly washed during aging with a salt solution. The different processes create different rind microbial communities.
It might be easier just to figure out which microbes are present on cheese rind, isolate them and study them in the lab. But Bonham and Dutton wanted to look at the microbial communities present on different cheeses, so to do that they used a technique they have utilized in other cheese rind studies — they scrape rind of different cheeses and bring them back to the lab. There, they run genetic analyses to see which microbes are present and who are members of the same community.
Dutton and her team had already found that aging similar types of cheese (bloomy, natural or washed)stored in similar environments — such as stringent humidity and temperatures — even if geographically distant, develop similar types of bacterial communities on the cheese's rind.
Now, the researchers have compared the genetic material of 165 cheese-associated bacteria to one another. They identified over 4,000 genes that were shared between the bacteria, including several large clusters of genes — that they called genomic islands — that were shared by many species by a process called horizontal gene transfer.

Representative colonies of bacteria from cheese analyzed in the study growing on a petri dish.

A graphic depiction of horizontal gene transfer analyzed in sample cheese bacteria, with the level of connection depicted through thickness of lines.

Representative colonies of bacteria from cheese analyzed in the study growing on a petri dish.

A graphic depiction of horizontal gene transfer analyzed in sample cheese bacteria, with the level of connection depicted through thickness of lines.
Horizontal gene transfer is the swapping of genetic material between bacteria present in the community at the same time. It has been identified as the mechanism that confers antibiotic resistance from resistant bacteria to nearby, previously non-resistant bacteria.
Three processes can be involved in horizontal gene transfer: transformation, transduction, and conjugation. In transformation, short fragments of naked DNA are taken up by other bacteria. Transductive transfer can occur when bacteriophages — viruses that infect bacteria — transfer DNA from one microbe into another during infection. DNA can also be transferred via the sexual pilus structure of some bacteria in a process called conjugation.
The researchers found all processes were active in horizontal gene transfer that occurred in cheese rind. As they write in the paper, "horizontal gene transfer has been studied for decades, but examining it in a more natural context is challenging because it requires studying an entire community of microbes, rather than studying them in isolation."
Identifying the functions of genes that are frequently transferred could help to identify the selective forces that are most important for adapting to the cheese rind environment. Horizontal gene transfer helps organisms acquire mechanisms to get nutrients with processes they didn't have, become pathogenic (disease-causing), or become resistant to antibiotics. The team found about 23% of the transferred genes in the cheese bacteria involved functions dealing with acquiring nutrients, especially iron.
Bacteria need iron to perform vital cellular functions, but iron is in short supply on the surface of cheeses. Bacteria often use specialized molecules called siderophores to scavenge for iron, then the iron-bound siderophores are taken back up into the cell. The new study found only the genes associated with the uptake systems were found in some of the shared gene clusters, revealing the importance of acquiring this vital function to the bacteria.
"This finding suggests that horizontal gene transfer has allowed some microbes to 'cheat' and take up iron-bound siderophores without expending energy to produce the siderophores themselves," the study authors write.
Now that the team has found a way to study horizontal gene transfer on cheese rind, and know that a multitude of genes are shared between the microbial community there, they plan to delve deeper in to how the transfer occurs, why, and what the results mean to the bacteria, cheese, and to us.
"Since horizontal gene transfer is prevalent in many microbial communities, including those important for human health, we're now trying to study how this process impacts microbial life and death in a community," Dutton said in a press release.
- Follow Invisiverse on Facebook and Twitter
- Follow WonderHowTo on Facebook, Twitter, Pinterest, and Google+
Cover image via PublicDomainPictures/Pixabay























Comments
Be the first, drop a comment!