Onshore, or on a boat, have you ever wondered what swims below in the dark water? Using standard equipment and a new process, marine scientists can now get a good look at what is swimming by—just by analyzing the water.
Taking stock of fish living or passing through a river, estuary, or other body of water is never easy. For years, fish stocks and migration patterns were understood only by trawling for fish with nets, a process that literally only captures a small portion of the action down below.
Without harming fish or using specialized equipment, researchers from the Rockefeller University Program for the Human Environment examined small samples of water to develop a wealth of information on fish species in the East and Hudson Rivers of New York.

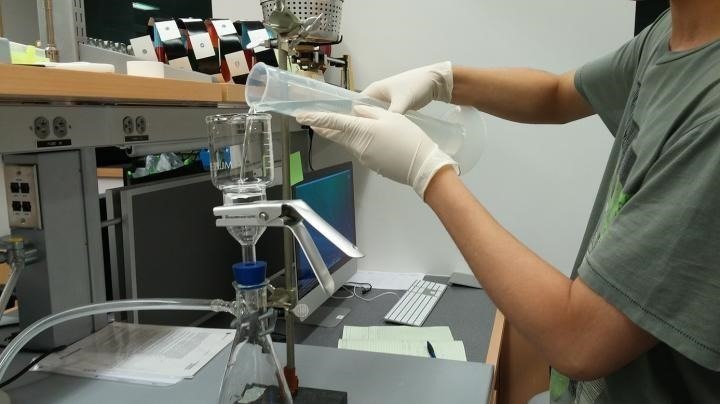
In a study published in PLOS ONE, scientists used environmental DNA, or eDNA, to evaluate fish populations over a six-month period, beginning in January 2016. Collecting 76 one-quart water samples at two environmentally different testing sites, the research team used genetic sequencing to identify microbial samples filtered out of the water.
In this study, environmental DNA refers to the invisible, microbial debris shed by fish as they swim through water. Whether molecules from the slime-coat that protects the skin of most fish, or from fecal matter deposited in the water, fish and other organisms leave a DNA trail in the water. Coauthor Zachary Charlop-Powers, a student researcher, noted in a press release that the analysis "uses the same methods that medical researchers employ to analyze human 'microbiomes.'"
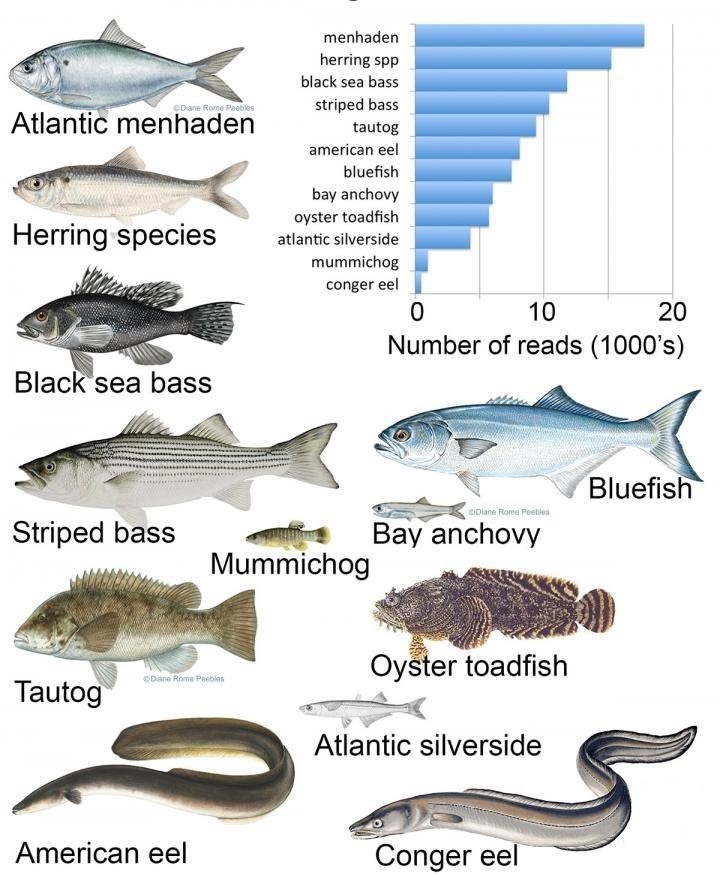
Once the eDNA samples are filtered and sequenced, scientists can identify fish species using the database GenBank, and surveys that were physically compiled between 1988 and 2015, among other resources. Each fish identification is called a "read." The frequency of the study reads, or DNA samples of particular fish, aligned with historical seasonal fish census data, underscoring the capability of the process for use in estimating fish stocks (or abundance).

In addition to offering a dynamic picture of the fish populations migrating or moving through the test waters, the research offered a wealth of information about other aspects of the marine environment, including:
- The eDNA analysis also captured microbial debris from domestic and wild animals, as well as humans.
- DNA from exotic fish not normally found in the test waters, like Atlantic salmon, European sea bass, and Pacific red snapper, are considered "menu fish." Their presence in the survey was attributed to their presence on a dinner plate, and later, their DNA passed through humans into raw sewage deposited into estuaries from which water samples were drawn.
- Overall, the analysis identified 42 fish species, including 81% of those known to be common local fish. On the reliability of the results, Senior Research Associate Mark Stoeckle said, "We didn't find anything shocking about the fish migration—the seasonal movements and the species we found are known already. It amazes me that we can get the same information from a small cup of water and a large net full of fish."
- Some of the DNA could not be traced to a specific fish species, a problem that will ultimately be resolved as DNA resource libraries grow.
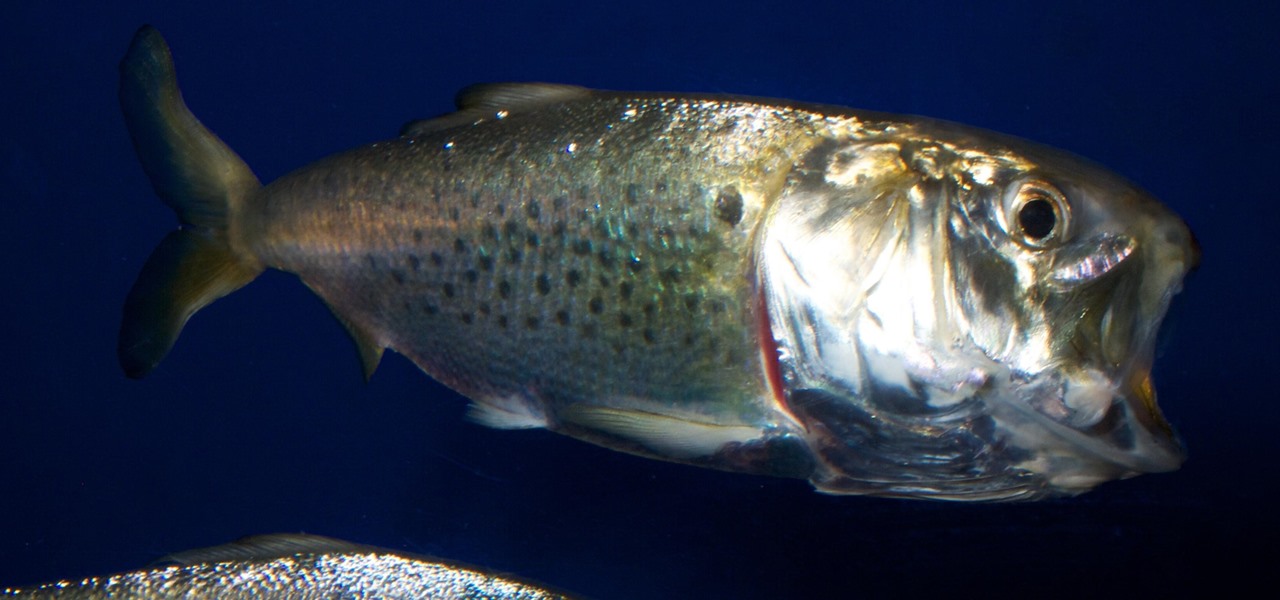
- If used to monitor fish species movements over time, the authors suggest an upswing in types of fish, like the Atlantic menhaden herring (pictured in this article's cover image), could explain the unlikely appearance of whales or dolphin in local New York waters.
Study authors predict the low-cost eDNA analysis process will be a mainstream tool in the next few years. From monitoring sentinel and other fish species without netting the fish, to identifying invasive species in ships' ballasts, or providing tourist boards with an understanding of what fish live in local lakes, collection of eDNA boosts data for future research.
Jesse Ausubel, Director of Program for the Human Environment, said in the press release, "Blood tests have now become so sensitive they can provide evidence of all kinds of conditions in a human body, so it is not really surprising that that we can now learn much more from tests for biological molecules circulating in water."
Just updated your iPhone? You'll find new emoji, enhanced security, podcast transcripts, Apple Cash virtual numbers, and other useful features. There are even new additions hidden within Safari. Find out what's new and changed on your iPhone with the iOS 17.4 update.




















Be the First to Comment
Share Your Thoughts